Input data and parameters
Input
| Analysis date: | Tue Oct 31 10:15:52 GMT 2023 |
| BAM file: | SRX7354383.markdup.sorted.bam |
| Counting algorithm: | uniquely-mapped-reads |
| GTF file: | gencode.v44.annotation_genes_collapsed_only_patched_ERCC92.gtf |
| Number of bases for 5'-3' bias computation: | 100 |
| Number of transcripts for 5'-3' bias computation: | 1,000 |
| Paired-end sequencing: | yes |
| Protocol: | strand-specific-reverse |
| Sorting performed: | yes |
Summary
Reads alignment
| Number of mapped reads (left/right): | 73,755,943 / 73,726,879 |
| Number of aligned pairs (without duplicates): | 73,717,107 |
| Total number of alignments: | 160,008,443 |
| Number of secondary alignments: | 12,525,621 |
| Number of non-unique alignments: | 20,488,376 |
| Aligned to genes: | 125,465,041 |
| Ambiguous alignments: | 525,162 |
| No feature assigned: | 12,416,909 |
| Missing chromosome in annotation: | 1,112,955 |
| Not aligned: | 3,607,772 |
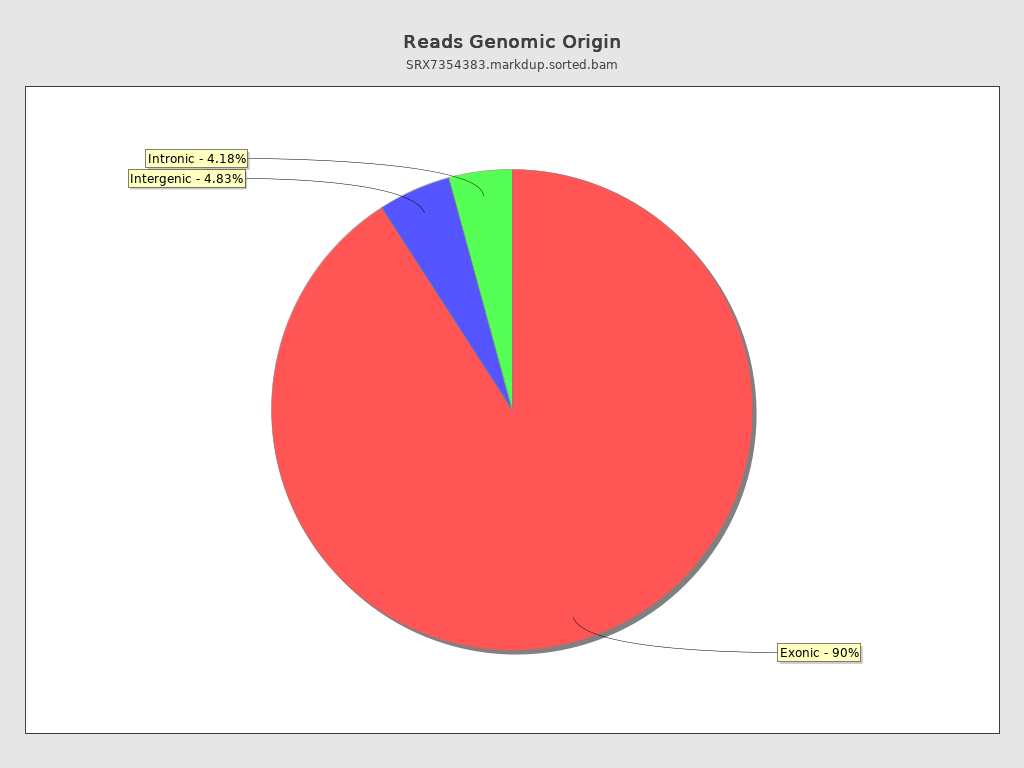
Reads genomic origin
| Exonic: | 125,465,041 / 90.99% |
| Intronic: | 5,756,691 / 4.18% |
| Intergenic: | 6,660,218 / 4.83% |
| Intronic/intergenic overlapping exon: | 5,399,234 / 3.92% |
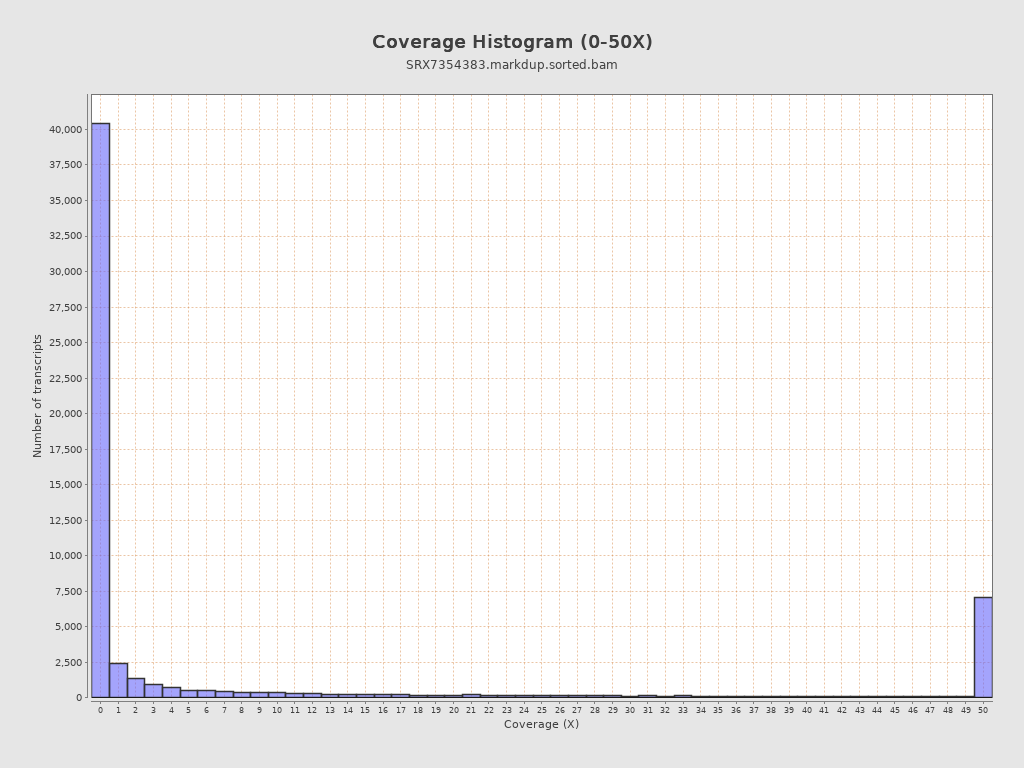
Transcript coverage profile
| 5' bias: | 0.14 |
| 3' bias: | 0.13 |
| 5'-3' bias: | 2.05 |
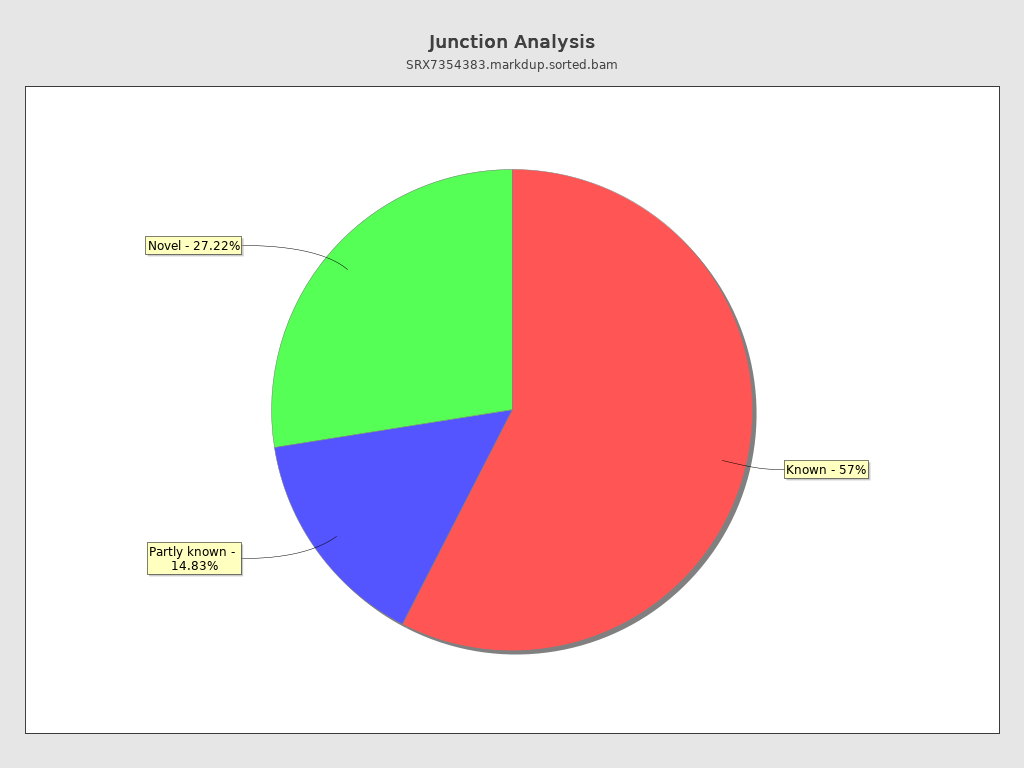
Junction analysis
| Reads at junctions: | 39,845,894 |
| ACCT | 5.21% |
| TCCT | 5.08% |
| ATCT | 4.49% |
| AGGT | 4.37% |
| CCCT | 3.26% |
| AGGA | 3.07% |
| GCCT | 2.65% |
| AGGC | 2.37% |
| TTCT | 2.06% |
| AGCT | 1.94% |
| TCCC | 1.72% |

.png)
.png)
.png)